Let’s start by writing the letter T using numpy in Python, encoded in a 0/1 array.
image_t = np.array( [[ 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0,], [ 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0,], [ 0, 0, 1, 1, 1, 1, 1, 1, 1, 1, 1, 0, 0,], [ 0, 0, 1, 1, 1, 1, 1, 1, 1, 1, 1, 0, 0,], [ 0, 0, 1, 1, 1, 1, 1, 1, 1, 1, 1, 0, 0,], [ 0, 0, 0, 0, 0, 1, 1, 1, 0, 0, 0, 0, 0,], [ 0, 0, 0, 0, 0, 1, 1, 1, 0, 0, 0, 0, 0,], [ 0, 0, 0, 0, 0, 1, 1, 1, 0, 0, 0, 0, 0,], [ 0, 0, 0, 0, 0, 1, 1, 1, 0, 0, 0, 0, 0,], [ 0, 0, 0, 0, 0, 1, 1, 1, 0, 0, 0, 0, 0,], [ 0, 0, 0, 0, 0, 1, 1, 1, 0, 0, 0, 0, 0,], [ 0, 0, 0, 0, 0, 1, 1, 1, 0, 0, 0, 0, 0,], [ 0, 0, 0, 0, 0, 1, 1, 1, 0, 0, 0, 0, 0,], [ 0, 0, 0, 0, 0, 1, 1, 1, 0, 0, 0, 0, 0,], [ 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0,], [ 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0,]])Let’s visualize this.
import matplotlib.pyplot as plt import matplotlib.cm as cm %matplotlib inline plt.imshow(image_t,cmap=cm.gray) plt.axis('off')This is what we get, indeed the array represents the letter “T”.
Now, let’s generate a corrupted version of this image by flipping each bit with some probability p. We can also do it a bit faster using utilities of numpy (corrupt_image_fast)
import random def corrupt_image(image,p=0.15): corrupted = image.copy() rows, cols = corrupted.shape for r in range(rows): for c in range(cols): if random.random() <= p : corrupted[r][c] = 1 - corrupted[r][c] #flip pixel return corrupted def corrupt_image_fast(image,p=0.15): corrupted = image.copy() rows, cols = corrupted.shape prob = np.random.rand(rows,cols) corrupted = np.multiply(corrupted, prob>p) + np.multiply(1-corrupted, prob<=p) return corruptedLet’s range the corruption probability from 0 to 1 with a step of 0.05 and see what we get.
plt.figure(1) index = 1 for p in np.linspace(0,1,21): plt.subplot(3,7,index) plt.title(repr("{:1.2f}".format(p))) plt.imshow( corrupt_image_fast(image_t,p), cmap=cm.gray) plt.axis('off') index+=1
Noisy version of the original image.
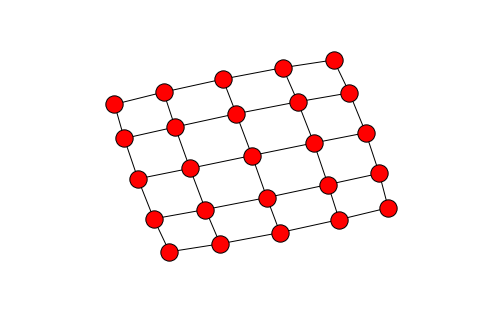
Therefore our problem informally is how can we get from the noisy image in the right to the original image? What are the modelling assumptions that we can use that would make the original image most likely? Think about other letters or digits. They are structured. The image consists of homogeneous patches that suddenly change when we get close to the boundaries. We impose the natural grid structure on the image, where each node corresponds to a pixel, and edges to neighboring pixels corresponding to the four possible directions, north, south, west, east. Here is a visualization of a
grid.
Modelling: Each node
on the grid has a binary value
, corresponding to whether the pixel is on or off in the image. We will model the recovery problem probabilistically. Let us assume that what we observe is the signal
and the original image is the signal
. We will maximize the conditional probability
over all possible original signals
. We impose our intuitive assumptions we discussed above by imposing an appropriate prior probability distribution on
. Specifically, we assume that the probability of
is proportional to
. Why is this a good prior? For any edge between two pixels
with the same value the exponent is equal to
. On the contrary, when the pixel values differ the exponent is equal to
. Hence the prior encourages homogeneous patches. This is known as a locally dependent Markov Random Field (MRF). We use Bayes’ rule to compute the conditional probability
we care about.
. First we compute the likelihood function
. Let
be the conditional density function for all
. For the sake of our exposition we have assumed in this lecture
. We will assume that the pixel noise is iid. Notice that there are real cases where the noise is localized and forms “patches” too. The iid assumption simplifies a lot the computation of the likelihood function as it factors over the pixels, namely
. According to Bayes’ rule
. Maximizing
is equivalent to maximizing the nominator as the denominator does not depend on
. This is convenient computationally as computing the marginal distribution
is not easy, at least the straightforward computation
requires summing over
possible values of
. Equivalently, due to the monotonicity of the
function we can try to maximize
.
The last term does not depend on our variable
, so our problem is to maximize the objective
where
. Observe that
may also take negative values. For example, suppose
. According to the probabilistic model we coded up in Python, if
,
. However, if
,
. Now we will refresh our combinatorial optimization toolbox. We are optimizing over discrete signals
, and there are
possible such signals since for each pixel there are two possible values. Observe that since
, and therefore
. This suggests that the second term has to do with a cut function since
is 1 if the edge
connects endpoints with different pixel values. It turns out that indeed our maximization problem can be reduced to a minimum cut/maximum flow computation. Specifically, we create a network as follows.
- We create two extra nodes, the source and the sink
respectively.
- We add an arc from the source
to each node
for which
. We set the capacity to be $\lambda_i$.
- We add an arc from each node
for which
to the sink $t$. We set the capacity to be $-\lambda_i$.
- For each pair of neighboring nodes
, we add two directed edges
with capacity
.
For any binary image let
, and
. The cut value is equal to
This is precisely our objective negated shifted by a constant. Hence, minimize this cut value is equivalent to our maximization problem. We can solve this using max flows in polynomial time! Here is an alternative view of our objective. We are given a binary matrix that is associated with a cost proportional to the number of edges of the corresponding grid that connect endpoints with different values (call such edges bad). We are allowed to flip an entry of the binary matrix if we pay dollars (i.e., per entry we change). For each bad edge we pay
dollars.Hence the total cost of our choice is equal to
This of course corresponds to a special case of the weights but captures fully our problem. Let’s see how we can translate all the above into Python code. We will use igraph, a useful package for graph algorithms in Python (another useful one is networkx) and for simplicity we will assume
for all neighboring
. Also let
according to our simple choice of a conditional density function. Let’s assume that
, therefore since
we get that
for all pixels such that
.
def create_graph(img, K=1, lambda=3<span id="mce_SELREST_start" style="overflow:hidden;line-height:0;"></span>.5): max_num = len(img)*len(img[0]) s,t = max_num, max_num + 1 edge_list = [] weights = [] for r_idx, row in enumerate(img): for idx, pixel in enumerate(row): px_id = (r_idx*len(row)) + idx #add edge to cell to the left if px_id!= 0: edge_list.append((px_id -1, px_id)) weights.append( K ) #add edge to cell to the right if px_id != len(row) -1: edge_list.append((px_id +1, px_id)) weights.append( K ) #add edge to cell to the above if r_idx!= 0: edge_list.append((px_id - len(row), px_id)) weights.append( K ) #add edge to cell to the below if r_idx != len(img) -1: edge_list.append((px_id + len(row), px_id)) weights.append( K ) #add an edge to either s (source) or t (sink) if pixel == 1: edge_list.append((s,px_id)) weights.append( lambda ) else: edge_list.append((px_id, t)) weights.append( lambda ) return edge_list, weights, s, tNow we can run max flow on this network to get our min cut as follows.
def recover(noisy, K=1, lambda=3.5): edge_list, weights, s, t = create_graph(noisy, K,lambda) g = igraph.Graph(edge_list) output = g.maxflow(s, t, weights) recovered = np.array(output.membership[:-2]).reshape(noisy.shape) return recovered plt.imshow(recover(corrupt_image_fast(image_t,0.15), 1, 3.5))For
we obtain the following recovery result for our image representing letter T.
For more, check out the following references:
[1] Lecture 1, Data Analytics class
[2] Exact Maximum a Posteriori inference for binary imagesPreliminaries
For those not familiar with Bayes’ rule and flows in networks, the Wikipedia articles are a good start.